New chromosome-level proof suggests Medicago evolution adopted a extra complicated, and stunning path, than beforehand thought.
A protracted-standing puzzle in plant evolution is beginning to come collectively. Scientists have used a strong imaging approach to hint how chromosome numbers modified in Medicago, a bunch that features alfalfa, one of the world’s most necessary forage crops. Their findings counsel that species with seven chromosomes did not gain complexity over time. Instead, they likely evolved by losing and reorganizing genetic material from an ancestor with eight chromosomes.
Instead of a single chromosome fusion, the transition involved extensive breakage and reassembly across three ancestral chromosomes. The study also highlights notable chromosome stability within the Medicago sativa complex, helping clarify links between genome structure, species formation, and crop evolution.
Challenges in Reconstructing Chromosome Evolution
Changes in chromosome number are common in plants, but determining the direction of those changes is often difficult because ancestral species are extinct or unknown. In Medicago, this question has remained unresolved for years. Scientists have debated whether species with seven chromosomes evolved from those with eight, or the other way around.
Although genome sequencing has advanced evolutionary studies, detailed chromosome-level evidence is still limited, especially in plants with repeat-rich and polyploid genomes. Standard cytogenetic methods often lack the resolution needed to follow entire chromosomes across related species. These challenges have made it difficult to map the evolutionary path of chromosome changes in Medicago.
Researchers from Shihezi University, Northwest A&F University, and Anhui Normal University reported in Horticulture Research that chromosome-specific painting can clarify the evolutionary history of karyotype changes in Medicago. Their findings suggest that species with seven chromosomes descended from an ancestor with eight through complex fission and fusion events. The study also confirms strong chromosome conservation among major alfalfa relatives.
Mapping Chromosome Rearrangements
The team developed a whole-chromosome painting system covering all eight chromosomes of alfalfa using more than 736,000 chromosome-specific oligonucleotides. They then used sequential fluorescence in situ hybridization to compare multiple Medicago accessions. This approach allowed direct visualization of chromosomes and comparisons between species with eight and seven chromosomes.
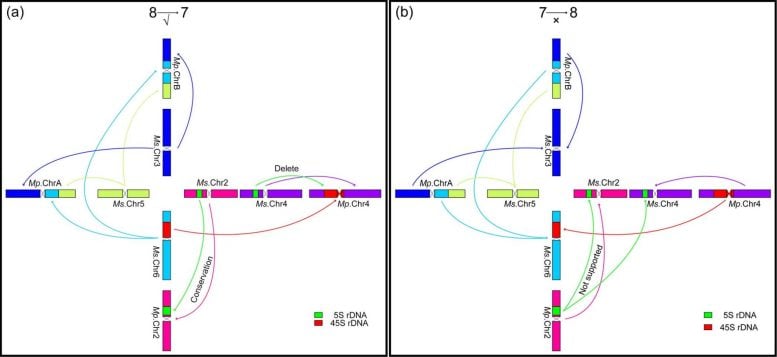
Within the M. sativa complex, including both diploid and autotetraploid forms, chromosome structures were highly stable, with no detectable rearrangements between chromosomes. In contrast, M. polymorpha showed a different pattern. Earlier theories suggested that its reduced chromosome number came from a simple fusion of chromosomes 3 and 7. However, the new evidence shows that chromosomes 3, 5, and 6 broke into segments and were reorganized into two new chromosomes.
Molecular Evidence Supports a Descending Model
The researchers also analyzed ribosomal DNA patterns. In species with x = 8, 5S rDNA appears on chromosomes 2 and 4, while 45S rDNA is located on chromosome 6. In M. polymorpha with x = 7, 5S rDNA remains only on chromosome 2, and 45S rDNA shifts to chromosome 4.
Together with the structural findings, these patterns strongly support a reduction from eight to seven chromosomes. Phylogenetic analysis suggests this transition occurred about 12 million years ago, while autotetraploidization in alfalfa likely happened around 6 million years ago.
The study shows that chromosome evolution in Medicago was not a simple one-step process. Instead, it involved multiple coordinated rearrangements. By combining cytogenetic observations with evolutionary analysis, the researchers provide clearer evidence of how chromosome changes contributed to diversification in this important crop group. The results also show that genome assemblies alone may not fully capture complex chromosome histories, especially in repeat-rich regions.
Implications for Breeding and Future Research
Beyond addressing an evolutionary question, the chromosome-painting method developed in this study has practical uses. It can help identify chromosomal differences among Medicago varieties, track karyotype stability over generations, and confirm chromosome inheritance in hybrids.
These applications make the technique valuable for both evolutionary research and crop improvement. More broadly, the study offers a useful framework for investigating chromosome evolution in other plant groups where similar genomic changes remain poorly understood.
Reference: “Chromosome-specific painting provides insights into the karyotype evolutionary direction and trajectory in the genus Medicago” by Wei Wang, Yuanbin Zhu, Xia Wu, Zixiang Guo, Qian Zheng, Guangzhen Shi, Yuanhao Li, Wenjun Luo, Fei Wang, Haitao Shen, Sheng Zuo, Quanliang Xie, Hongbin Li and Zhuang Meng, 14 November 2025, Horticulture Research.
DOI: 10.1093/hr/uhaf313
The research was supported by the Natural Science Foundation of China (grant number 32400496), Tianchi Talent Project of Xinjiang (grant number CZ001604), High-level Talents Scientific Startup Project of Shihezi University (grant number RCZK202362), Basic Research General Project under the Science and Technology Program of Xinjiang Production and Construction Corps (grant number 2024DA021, 2024DA030), Shihezi University Youth Top-notch Training Project (grant number BJZK202403), Science and Technology Research Project of Bingtuan (grant number 2023AB070, 2025AB084), and Science and Technology Innovation Talent Project of Bingtuan (grant number 23CB008-01).
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.