
Proteins — massive molecules with a particular organic operate — are created by stringing collectively smaller molecular constructing blocks (amino acids) into chains which fold into advanced 3D shapes. Credit Johan Jarnestad/The Royal Swedish Academy of Sciences
The variety of recognized proteins is infinitely small compared to the universe of potential proteins which might in principle be realized. Yet these recognized proteins are the one main coaching floor for future protein design. Understanding how consultant these proteins are of the general potential variety can due to this fact assist inform methods for a variety of functions, together with therapeutic, biocatalysis, or biomaterials growth.
Published in PNAS, a world staff from the Okinawa Institute of Science and Technology (OIST), the Institute of Science and Technology Austria (ISTA), the University of Vienna and the Centro de Astrobiología (CAB) investigated the connection between protein evolution and sequence house, figuring out the limiting components behind protein diversification. Their findings reinforce theories of DNA recombination as a driving drive of ancestral protein formation and spotlight the constraints of many cutting-edge AI protein design strategies.
“Modern AI methods are thought to be revolutionizing protein design, with the 2024 Nobel Prize in Chemistry awarded to the team behind AlphaFold. Yet most of these AI design methods are typically trained on databases of known proteins. So without understanding how representative these known proteins are of sequence space, how confident can we be that such methods can generate truly diverse protein designs?” says Professor Fyodor Kondrashov, head of OIST’s Evolutionary and Synthetic Biology Unit.
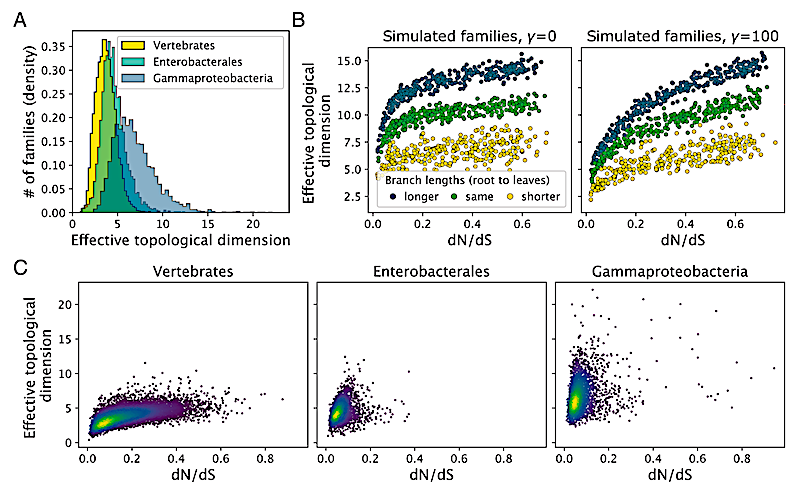
Dimensionality of pure and simulated sequences. (A) The distribution of the efficient topological dimension in numerous protein households. (B) Dimensionality of a simulated protein household as a operate of imply dN/dS per household (an empirical estimate of α, the fraction of allowed amino acid states), and department lengths (colour). (C) Estimated dimensionality as a operate of imply dN/dS per household for pure sequences. Brighter colours (extra yellow) point out larger level density. — PNAS
Imagine you could have 20 or so totally different block varieties, which you’ll be able to join in numerous orders and abundances into chains of tens, tons of and even hundreds of blocks in size. Mapping all potential ensuing chains creates a sequence house.
For proteins, the form and construction of their amino acid constructing blocks imply solely a minute fraction of potential protein sequences can fold up into the proper 3D form to energy a organic operate. They want the proper chemical teams within the appropriate locations to create the interactions that can keep 3D form or bind to different molecules. Mapping the sequences that fulfil this requirement creates a smaller useful house.
Of these potential useful sequences, it’s doubtless that comparatively few have ever existed throughout evolutionary historical past. Therefore, the researchers got down to uncover how consultant this subset of proteins is of useful house.
The researchers began by mathematically describing the sequence house taken up by recognized proteins. They then constructed a mannequin of protein evolutions to know the organic components controlling the structural diversification of a variety of naturally-occurring protein households. From their fashions, they then predicted what number of useful sequences they’d anticipate to exist for a given organic operate.
By evaluating the range of recognized proteins to those theoretical predictions of protein evolution, the researchers discovered that point-of-origin results far outweighed the affect of different key evolutionary processes.
“That starting point is the main evolutionary limit is not necessarily surprising, but the scale of its importance is really quite remarkable,” observes lead creator Lada Isakova, PhD scholar throughout the unit. “As an evolutionary biologist, I was intrigued to see how little selection and epistasis seemed to matter in our results.”
What limits protein evolution?
When mutations come up within the genes encoding for a selected protein, these can lead to adjustments to the sequence of amino acids produced, inflicting protein evolution. Natural choice limits which mutations persist over time primarily based on whether or not they enhance or hurt the protein’s operate or stability. Epistasis — genetic interactions leading to totally different outputs — additionally constrains evolution, as mutations could have restricted results alone, however massive results when current together with sure different mutations.
Both choice and epistasis are recognized to affect protein evolution, but Isakova and colleagues discovered that by far, the limiting issue of protein variety is the origins of our proteins, with comparatively small divergence seen from the areas of sequence house of ancestral proteins.
This analysis gives new insights into the origins of life, reinforcing current theories on preliminary protein formation. Isakova explains, “Our simulations suggest that, for the first proteins in the last universal common ancestor to arise, they couldn’t just diverge from mutations of a single first sequence, given the time constraints we see. Instead, small pieces of DNA must have shuffled around and recombined to create new DNA molecules which could encode very different proteins.”
The staff additionally hopes that the analysis conjures up experimental scientists to broaden the recognized sequence house. Isakova feedback, “Neural network approaches for functional protein prediction are limited by the data sets we provide. So based on existing data, most methods won’t be able to generalize well beyond the current known sequence space. We can see there’s huge swaths of sequence space left to be explored, but it’ll take new experimental data to enable expansion into these unknown realms.”
This international collaboration was supported by a Japan Science and Technology Agency (JST) Adopting Sustainable Partnerships for Innovative Research Ecosystem (ASPIRE) grant, which goals to construct a community between high researchers in Japan and all over the world, nurturing future scientific leaders.
Descent from a common ancestor restricts exploration of protein sequence space, PNAS (open entry)
Astrobiology,
